Note
Click here to download the full example code
Data generation mechanism¶
This example illustrates the Geometric SMOTE data generation mechanism and the usage of its hyperparameters.
# Author: Georgios Douzas <gdouzas@icloud.com>
# Licence: MIT
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import make_blobs
from imblearn.over_sampling import SMOTE
from gsmote import GeometricSMOTE
print(__doc__)
XLIM, YLIM = [-3.0, 3.0], [0.0, 4.0]
RANDOM_STATE = 5
def generate_imbalanced_data(
n_maj_samples, n_min_samples, centers, cluster_std, *min_point
):
"""Generate imbalanced data."""
X_neg, _ = make_blobs(
n_samples=n_maj_samples,
centers=centers,
cluster_std=cluster_std,
random_state=RANDOM_STATE,
)
X_pos = np.array(min_point)
X = np.vstack([X_neg, X_pos])
y_pos = np.zeros(X_neg.shape[0], dtype=np.int8)
y_neg = np.ones(n_min_samples, dtype=np.int8)
y = np.hstack([y_pos, y_neg])
return X, y
def plot_scatter(X, y, title):
"""Function to plot some data as a scatter plot."""
plt.figure()
plt.scatter(X[y == 1, 0], X[y == 1, 1], label='Positive Class')
plt.scatter(X[y == 0, 0], X[y == 0, 1], label='Negative Class')
plt.xlim(*XLIM)
plt.ylim(*YLIM)
plt.gca().set_aspect('equal', adjustable='box')
plt.legend()
plt.title(title)
def plot_hyperparameters(oversampler, X, y, param, vals, n_subplots):
"""Function to plot resampled data for various
values of a geometric hyperparameter."""
n_rows = n_subplots[0]
fig, ax_arr = plt.subplots(*n_subplots, figsize=(15, 7 if n_rows > 1 else 3.5))
if n_rows > 1:
ax_arr = [ax for axs in ax_arr for ax in axs]
for ax, val in zip(ax_arr, vals):
oversampler.set_params(**{param: val})
X_res, y_res = oversampler.fit_resample(X, y)
ax.scatter(X_res[y_res == 1, 0], X_res[y_res == 1, 1], label='Positive Class')
ax.scatter(X_res[y_res == 0, 0], X_res[y_res == 0, 1], label='Negative Class')
ax.set_title(f'{val}')
ax.set_xlim(*XLIM)
ax.set_ylim(*YLIM)
def plot_comparison(oversamplers, X, y):
"""Function to compare SMOTE and Geometric SMOTE
generation of noisy samples."""
fig, ax_arr = plt.subplots(1, 2, figsize=(15, 5))
for ax, (name, ovs) in zip(ax_arr, oversamplers):
X_res, y_res = ovs.fit_resample(X, y)
ax.scatter(X_res[y_res == 1, 0], X_res[y_res == 1, 1], label='Positive Class')
ax.scatter(X_res[y_res == 0, 0], X_res[y_res == 0, 1], label='Negative Class')
ax.set_title(name)
ax.set_xlim(*XLIM)
ax.set_ylim(*YLIM)
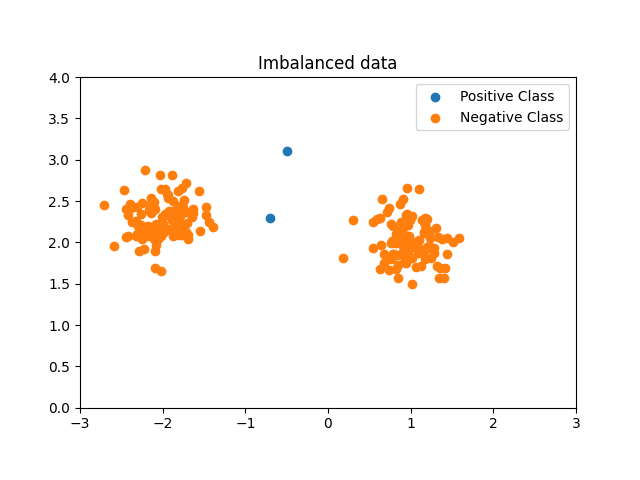
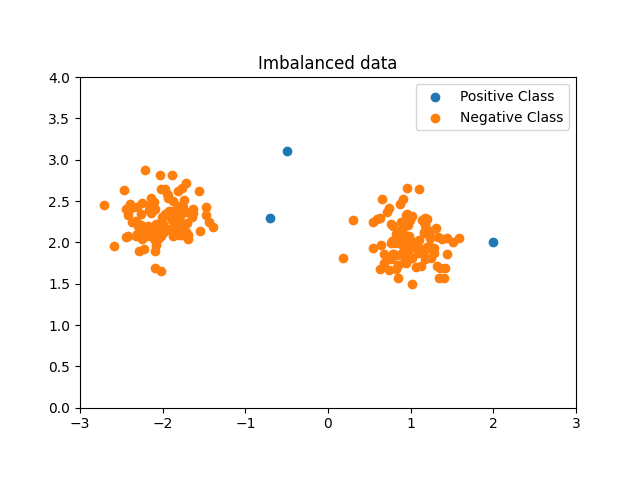
Generate imbalanced data¶
We are generating a highly imbalanced non Gaussian data set. Only two samples from the minority (positive) class are included to illustrate the Geometric SMOTE data generation mechanism.
Geometric hyperparameters¶
Similarly to SMOTE and its variations, Geometric SMOTE uses the k_neighbors
hyperparameter to select a random neighbor among the k nearest neighbors of a
minority class instance. On the other hand, Geometric SMOTE expands the data
generation area from the line segment of the SMOTE mechanism to a hypersphere
that can be truncated and deformed. The characteristics of the above geometric
area are determined by the hyperparameters truncation_factor,
deformation_factor and selection_strategy. These are called geometric
hyperparameters and allow the generation of diverse synthetic data as shown
below.
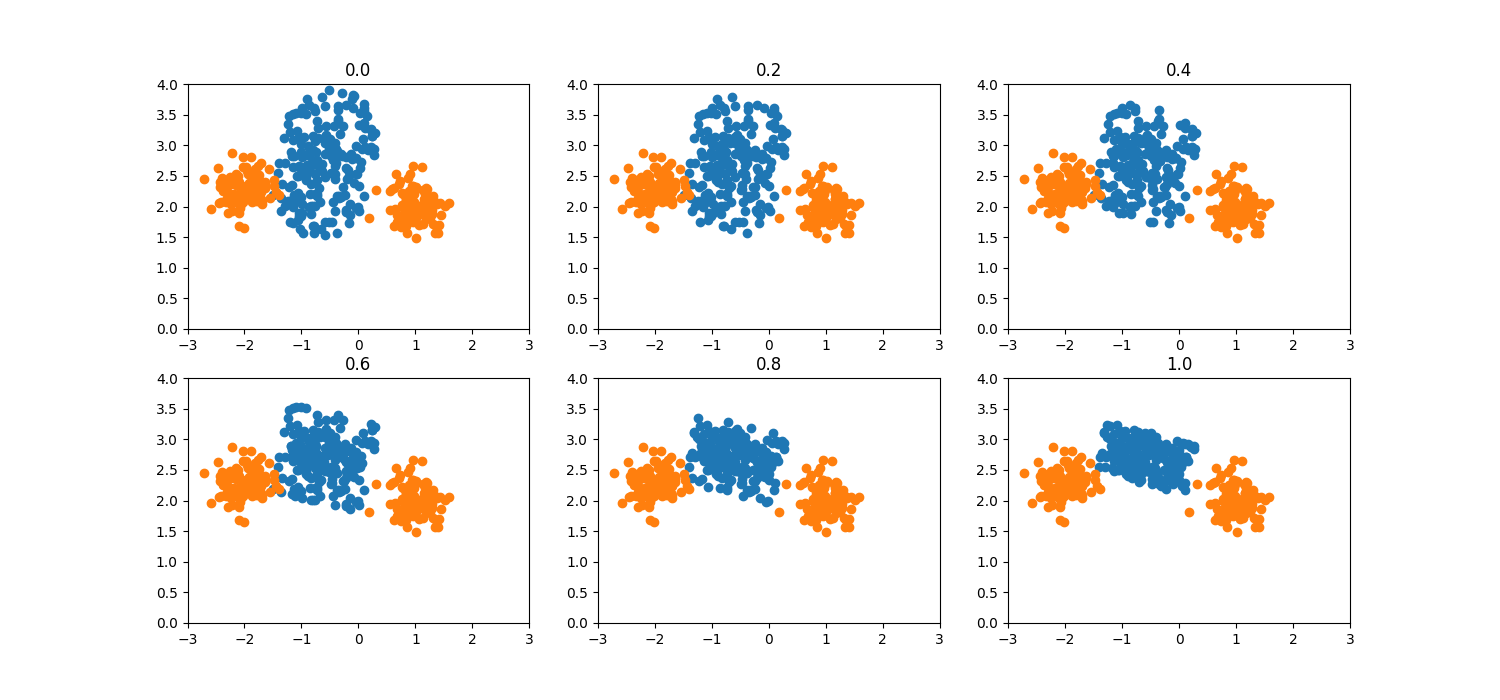
Truncation factor¶
The hyperparameter truncation_factor determines the degree of truncation
that is applied on the initial geometric area. Selecting the values of
geometric hyperparameters as truncation_factor=0.0,
deformation_factor=0.0 and selection_strategy='minority', the data
generation area in 2D corresponds to a circle with center as one of the two
minority class samples and radius equal to the distance between them. In the
multi-dimensional case the corresponding area is a hypersphere. When
truncation factor is increased, the hypersphere is truncated and for
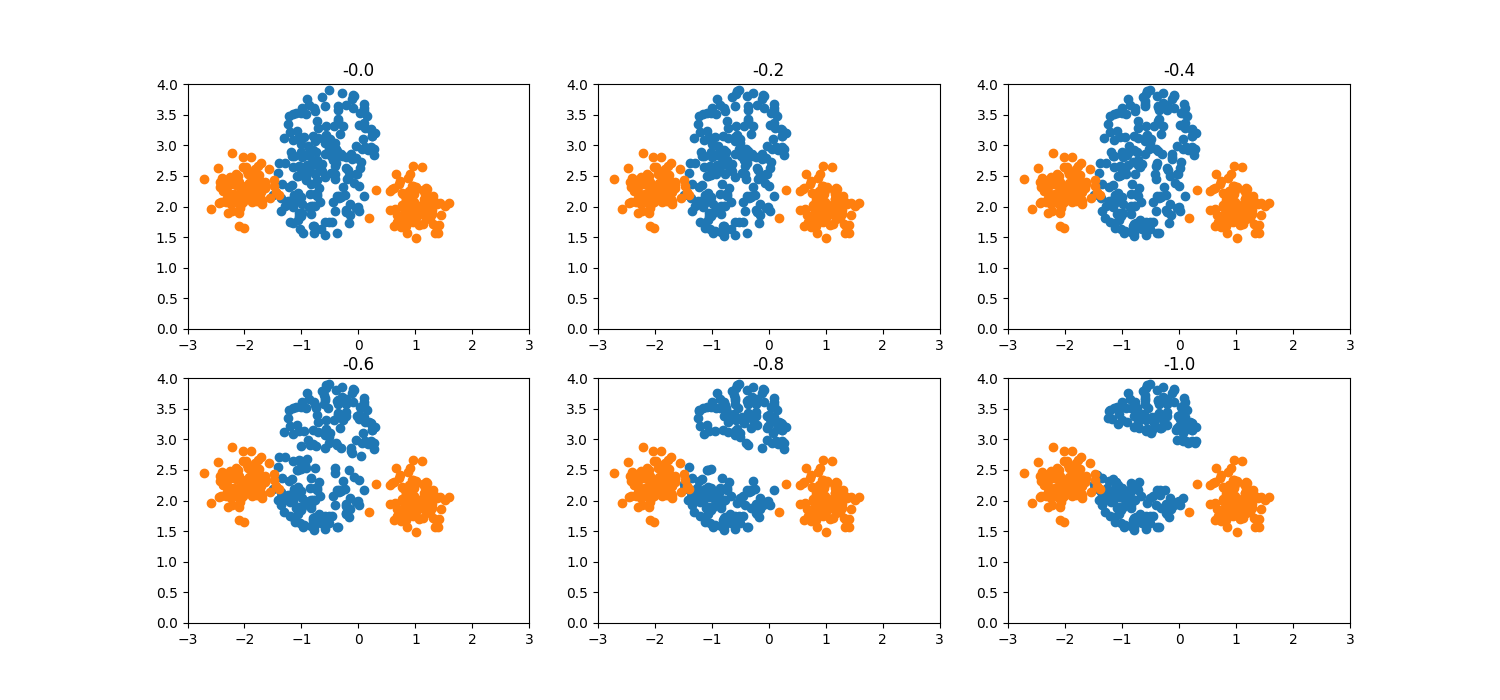
truncation_factor=1.0 becomes a half-hypersphere. Negative values of
truncation_factor have a similar effect but on the opposite direction.
gsmote = GeometricSMOTE(
k_neighbors=1,
deformation_factor=0.0,
selection_strategy='minority',
random_state=RANDOM_STATE,
)
truncation_factors = np.array([0.0, 0.2, 0.4, 0.6, 0.8, 1.0])
n_subplots = [2, 3]
plot_hyperparameters(gsmote, X, y, 'truncation_factor', truncation_factors, n_subplots)
plot_hyperparameters(gsmote, X, y, 'truncation_factor', -truncation_factors, n_subplots)
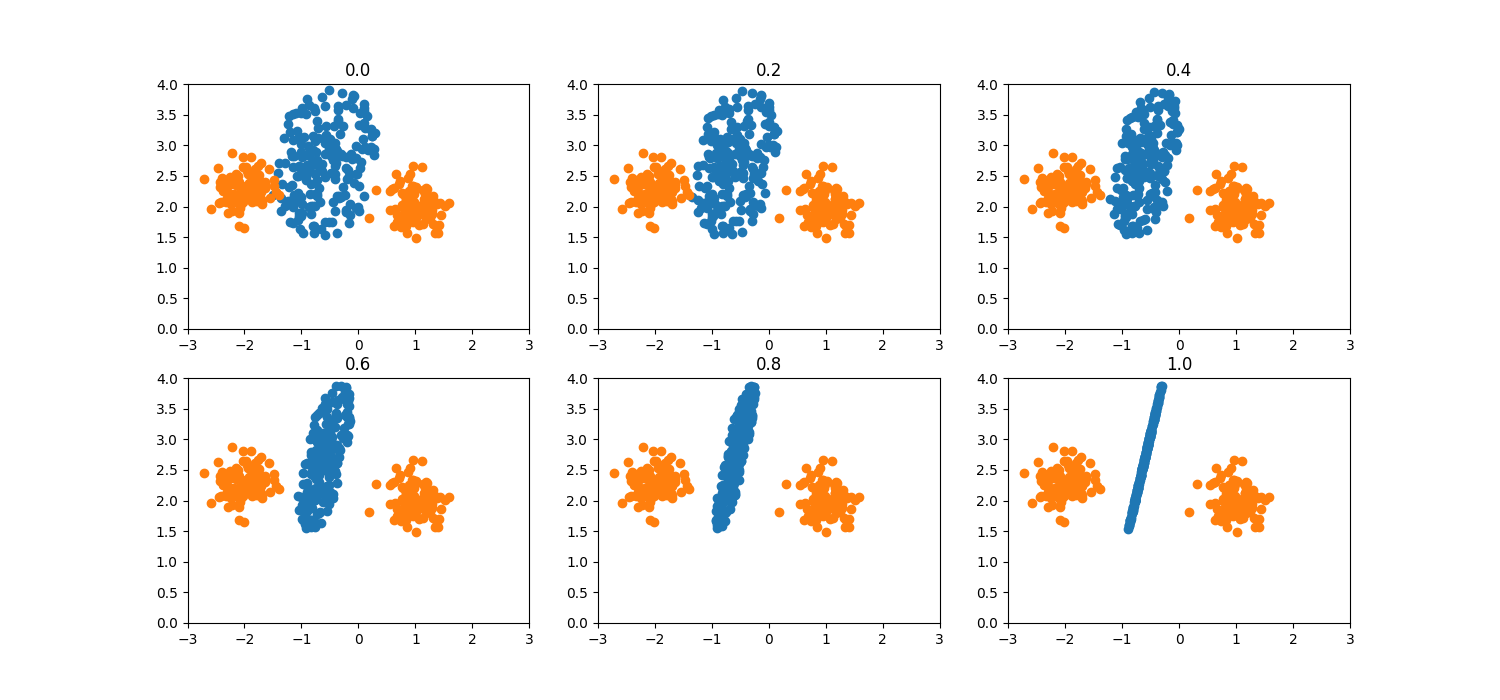
Deformation factor¶
When the deformation_factor is increased, the data generation area deforms
to an ellipsis and for deformation_factor=1.0 becomes a line segment.
gsmote = GeometricSMOTE(
k_neighbors=1,
truncation_factor=0.0,
selection_strategy='minority',
random_state=RANDOM_STATE,
)
deformation_factors = np.array([0.0, 0.2, 0.4, 0.6, 0.8, 1.0])
n_subplots = [2, 3]
plot_hyperparameters(gsmote, X, y, 'deformation_factor', truncation_factors, n_subplots)
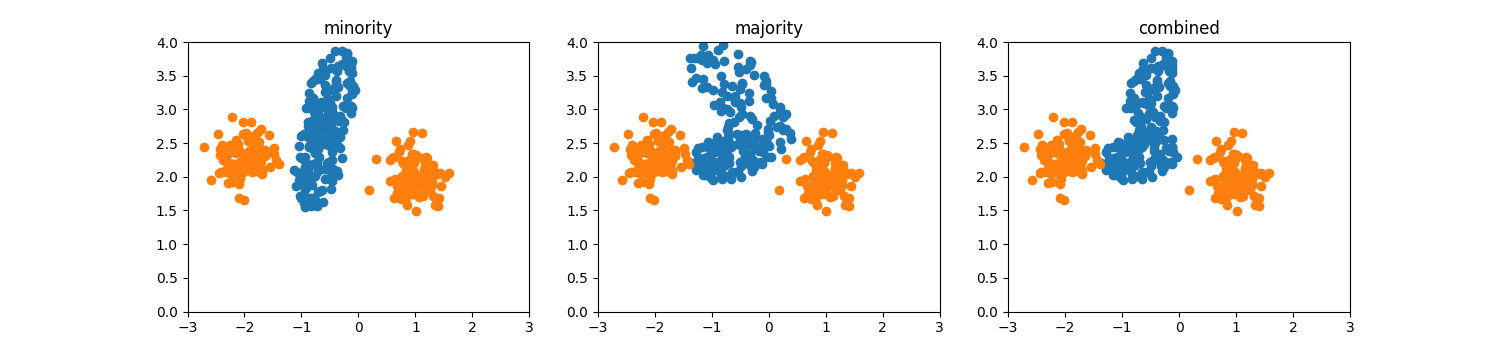
Selection strategy¶
The hyperparameter selection_strategy determines the selection mechanism
of nearest neighbors. Initially, a minority class sample is selected randomly.
When selection_strategy='minority', a second minority class sample is
selected as one of the k nearest neighbors of it. For
selection_strategy='majority', the second sample is its nearest majority
class neighbor. Finally, for selection_strategy='combined' the two
selection mechanisms are combined and the second sample is the nearest to the
first between the two samples defined above.
gsmote = GeometricSMOTE(
k_neighbors=1,
truncation_factor=0.0,
deformation_factor=0.5,
random_state=RANDOM_STATE,
)
selection_strategies = np.array(['minority', 'majority', 'combined'])
n_subplots = [1, 3]
plot_hyperparameters(
gsmote, X, y, 'selection_strategy', selection_strategies, n_subplots
)
Noisy samples¶
We are adding a third minority class sample to illustrate the difference between SMOTE and Geometric SMOTE data generation mechanisms.
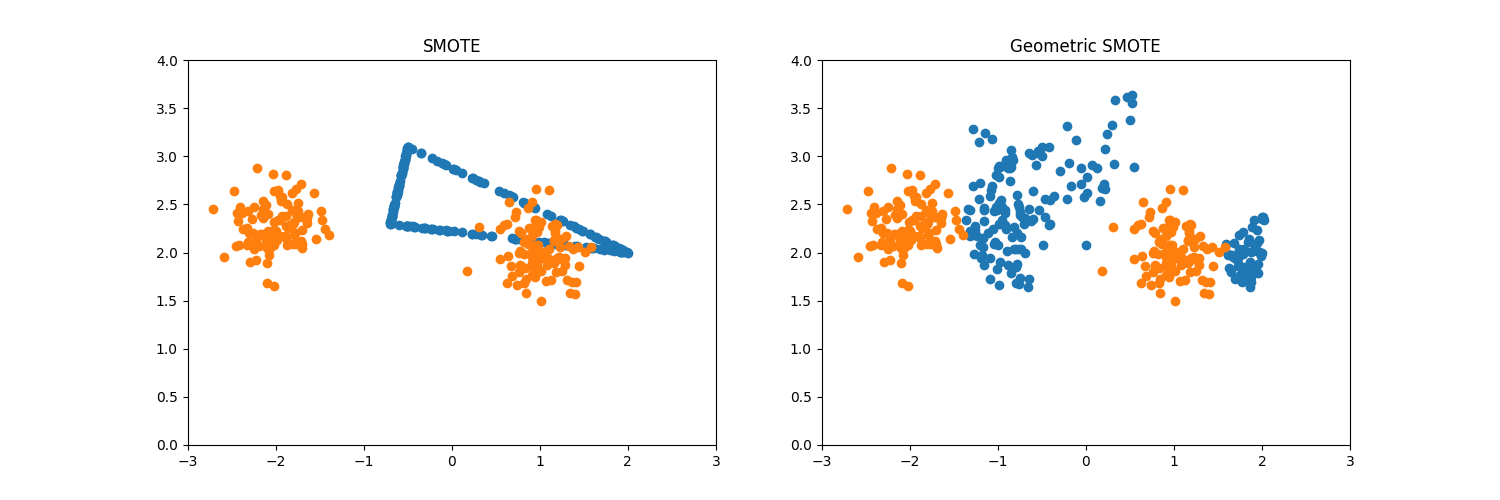
When the number of k_neighbors is increased, SMOTE results to the
generation of noisy samples. On the other hand, Geometric SMOTE avoids this
scenario when the selection_strategy values are either combined or
majority.
oversamplers = [
('SMOTE', SMOTE(k_neighbors=2, random_state=RANDOM_STATE)),
(
'Geometric SMOTE',
GeometricSMOTE(
k_neighbors=2, selection_strategy='combined', random_state=RANDOM_STATE
),
),
]
plot_comparison(oversamplers, X_new, y_new)
Total running time of the script: ( 0 minutes 2.346 seconds)